I'm building an app that listens to a doctor talk to a patient and writes the clinical note for them. The whole thing runs on a laptop. No internet, no cloud, no audio leaving the device. Two AI models do the work. One hears audio and types out words (like a really fast transcriptionist), and the other reads that transcript and writes the actual clinical note.
Something Felt Off
Before I ran my own evaluation, a teammate had evaluated 7 models using Gemini 3 Pro as the judge: 24 transcripts, 2 prompt variants, 7 rubric criteria, 336 scored results. Gemma 3n scored 77%, only 6 points behind the cloud model. The conclusion was "local models are surprisingly viable."
That's the eval I used to pick Gemma 3n. It got a lot right. But as I built the app and started reading the notes on complex visits, important details were missing. I'd read the transcript, then read the generated note, and think: where did half this visit go?
I went back to the heatmap and found the clue hiding in plain sight:
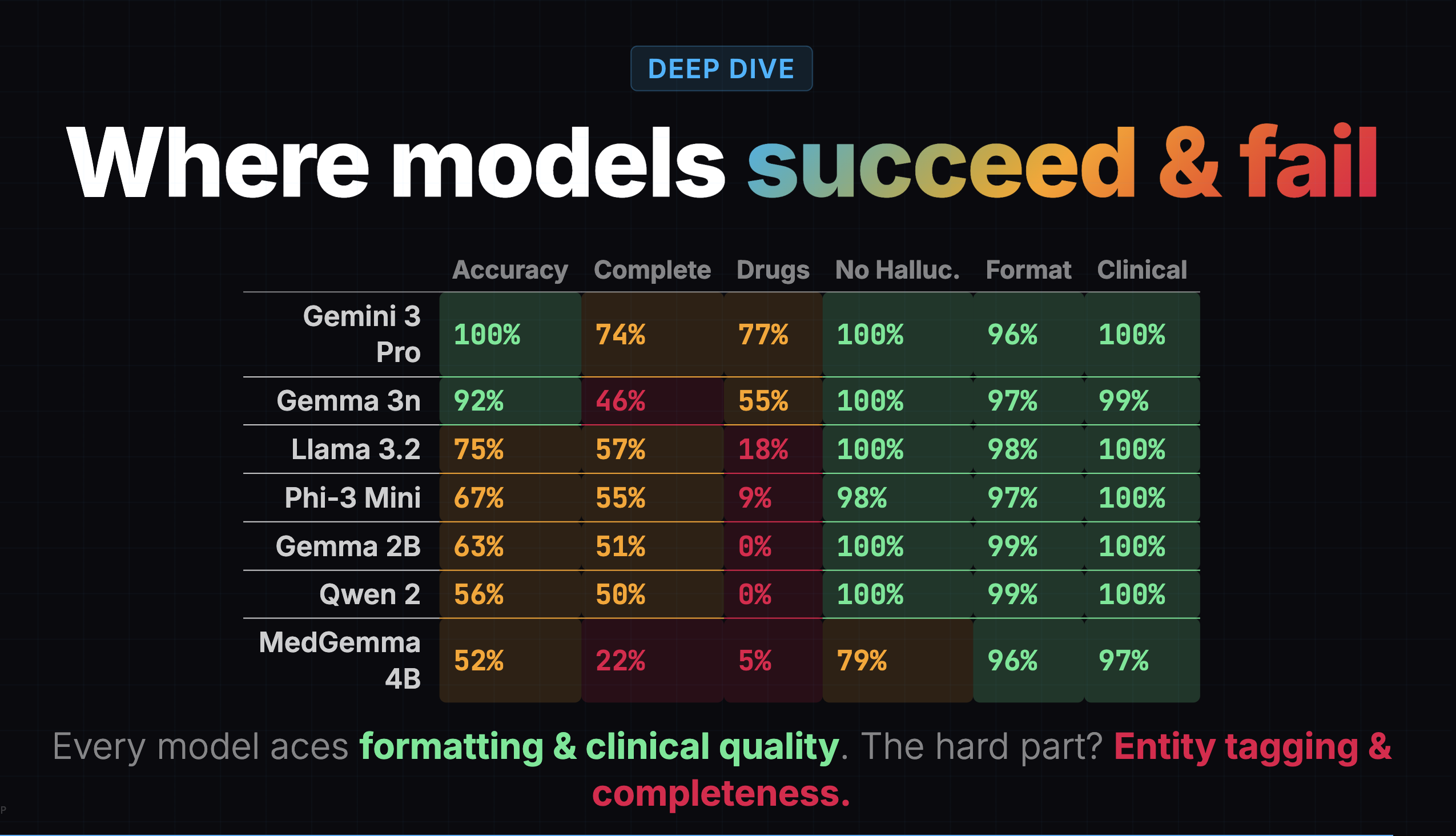

Gemma 3n's completeness score was 46%. It was capturing less than half the clinical content. But the overall score of 77% buried this because formatting (97%), clinical quality (99%), and hallucination resistance (100%) were all near-perfect.
A confident-looking note with missing content is arguably worse than a messy note with everything in it, because the doctor might not realize something is missing.
Grading My Own Grading
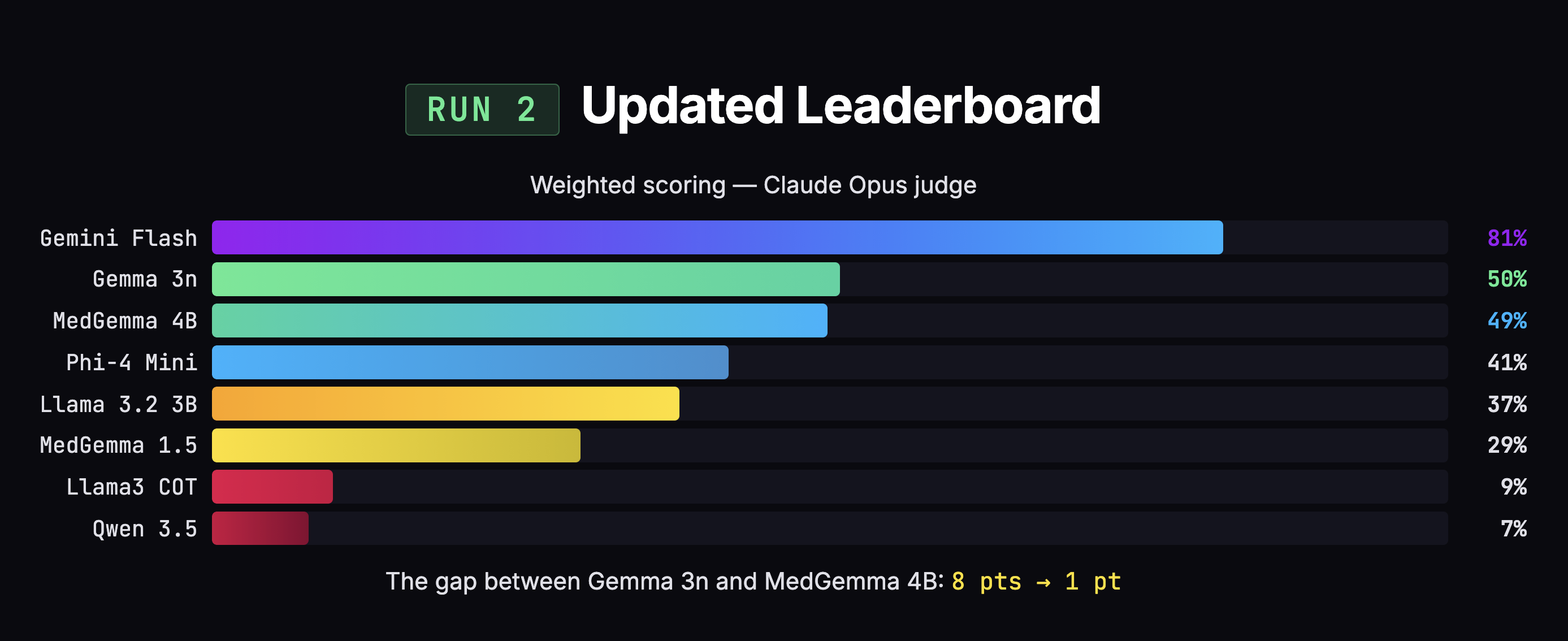
I ran my own eval: 8 models, 25 transcripts, 200 SOAP notes, two AI judges. The initial results said Gemma was beating MedGemma by 8 points, 49% to 41%. Case closed, right?
Not quite. I found three problems with my own scoring.
1. Double-penalization. When a model put content in the wrong section, the judge docked points for both "you missed content" and "you put content in the wrong place." Same information, penalized twice. This disproportionately hurt MedGemma, which captures more information but organizes it messily.
2. Clinical boilerplate flagged as hallucination. Phrases like "alert and oriented" and "no known drug allergies" show up in virtually every real clinical note. But because the doctor didn't explicitly say them, the judge flagged them as fabricated. That's like penalizing a note-taker for writing "Meeting started at 2 PM" when nobody announced the time out loud.
3. Equal weighting. Entity marking was broken for every local model (0-3%), making it dead weight that dragged down scores equally. Meanwhile, hallucination and completeness carried the same weight as formatting. A doctor can fix bad formatting in two seconds. A doctor cannot fix missing information without re-listening to the recording.
After fixing those three issues and re-scoring all 200 notes, the 8-point gap collapsed to 1 point (50% vs 49%). MedGemma jumped +8 points. Gemma moved +1.
The most striking example: MedGemma 4B. In the earlier eval: 53% overall, dead last, 22% completeness, 79% hallucination (worst of any model). In my eval: 49% overall, 48% completeness, 53% hallucination. Same model, dramatically different picture. The difference wasn't the model. It was the rubric.
MedGemma went from worst to essentially first among local models not because anything changed about the model, but because the scoring system went from ambiguous to explicit on the edge cases that affected it most.
Choosing by Failure Mode
The overall scores were too close to decide on numbers alone. So I looked at how each model fails. On the dimension that matters most, completeness, MedGemma grabbed 48% of the information versus Gemma's 35%.
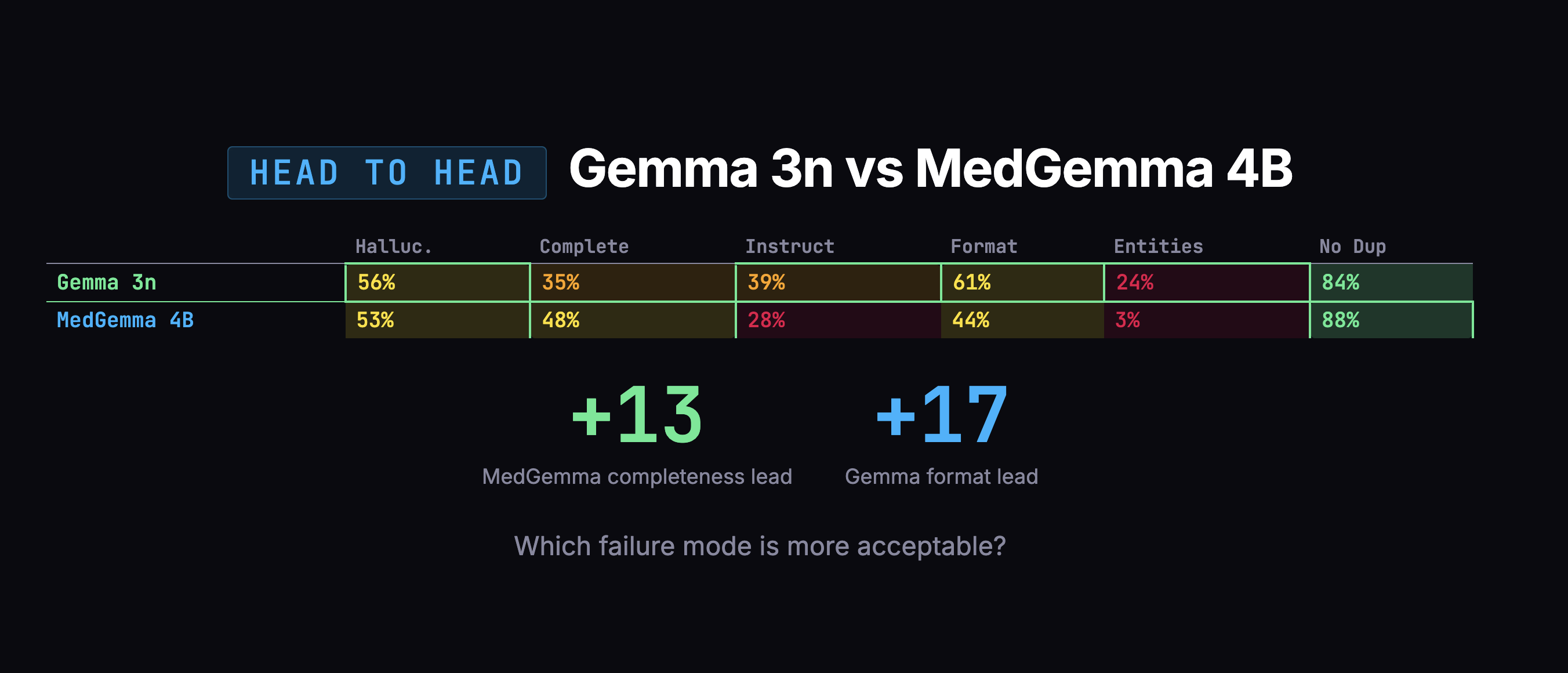

Gemma's failures were scarier: confusing which family member had a medical condition, duplicating entire notes. MedGemma's failures were mostly messy organization and verbosity, annoying, but fixable in post-processing.
I swapped engines.
There was one more thing I had to fix, and it was my own fault. I'd been telling all these models to format note headings using bold text (**Subjective** instead of # Subjective). Every AI model was trained on text where # means a heading. Asking a small model to use bold text as a heading is like asking someone fluent in English to write in a dialect they've never seen. Once I switched to standard markdown headings, I removed a whole category of unnecessary errors, and more importantly, removed what would have been a tax on every future fine-tuning run.
Picking the Voice Model
The popular choice for voice-to-text is Whisper. It works well for general speech. But doctors say things like "amlodipine" and "bilateral crackles." Whisper hears "am low to peen" and "by lateral crackles."
Google released MedASR through their Health AI Developer Foundations program. The numbers made this easy:
- Word Error Rate: MedASR ~5% on medical words, Whisper 25-32%.
- Model Size: MedASR 105M parameters, Whisper 1.5B (15x larger).
- Training Data: MedASR was trained on 5,000 hours of clinical audio. Whisper was trained on general internet speech.
Not a close call.
Then I hit a bug. A doctor says "thirty year old male patient." The app transcribes it as "three year old male patient." The problem: greedy decoding. MedASR processes audio in tiny slices and picks the best-matching word for each slice in isolation, with no memory of previous picks and no awareness of what's coming next. The word "three" sounds almost identical to the beginning of "thirty," but "three" exists as a single token in MedASR's vocabulary while "thirty" has to be assembled from four pieces (th + ir + t + y). By the time the "-ty" arrives, the model has already committed.
Google ships a 672 MB language model alongside MedASR. It was already on my machine. I just wasn't using it. Once plugged in, the model considers 8 possible interpretations simultaneously instead of committing one at a time. The language model knows "thirty year old" is far more probable than "three year old" in a clinical context.
There was a second issue: corrupted audio. MedASR expects clean audio at 16,000 samples per second. The app's browser environment captures at 48,000 and was downsampling badly, just picking every third sample and discarding the rest. On top of that, browser features designed for video calls (echo cancellation, noise suppression, auto gain) were processing the audio in ways MedASR wasn't trained on. Letting the browser handle sample rate conversion natively and disabling all video-call processing, combined with the language model fix, cut word errors by roughly 30%.
Fine-Tuning MedGemma
Fine-tuning takes a pre-trained model and shows it examples of what "good" looks like until it learns the pattern. I needed doctor-patient transcripts paired with high-quality clinical notes.
The transcripts came from MTS-Dialog, a dataset of 492 medical conversations. I selected 200, balanced across 19 specialties. For the notes, I used knowledge distillation: Gemini 2.5 Pro as a "teacher" generating ideal notes for each transcript across 7 templates (SOAP, H&P, Progress Note, Discharge Summary, Procedure Note, Consultation Note, DAP). That produced 1,394 usable training examples.
One discovery proved important: the training data and the live app must format instructions identically. If the training data says "Generate a SOAP note" but the live app sends the full template with section headers and formatting rules, the model sees unfamiliar input and performance drops. I caught this early and aligned both formats.
I split by conversation, not by note: all 7 notes from the same transcript go into the same group. Otherwise the model could train on conversation #42's SOAP note and then get "validated" on its DAP note, already knowing the patient.
Training ran on Modal, a cloud GPU platform, using an NVIDIA A100 with 80GB of memory. I used LoRA (Low-Rank Adaptation), which freezes the base model and attaches small trainable layers alongside it. Think of it like putting a thin lens in front of a camera rather than rebuilding the entire camera. I trained just 38 million parameters, roughly 1% of the model.
Two Runs, One Tradeoff
Run A also trained the model's embedding layer and output head, the layers responsible for converting words into numbers (input) and numbers back into words (output). Trainable parameters jumped from 38 million (1%) to 1.38 billion (24%). It started learning entity tagging, wrapping medications and conditions in structured markup like {{drug:metformin}} so the app can highlight them. But it overfitted fast:
| Step | Training Loss | Validation Loss | Status |
|---|---|---|---|
| 25 | - | 0.578 | Improving |
| 50 | 0.444 | 0.479 | Best checkpoint |
| 75 | 0.346 | 0.512 | Getting worse |
| 100 | 0.206 | 0.666 | Memorizing |
| 150 | 0.083 | 0.891 | Fully memorized |
Training loss measures how well the model performs on examples it has already seen. Validation loss measures performance on held-out examples it hasn't seen. At step 50, they're close: 0.444 vs 0.479. That's the sweet spot. After step 50, the gap blows open: training loss plummets to 0.083 while validation loss climbs to 0.891. The model is getting better at reciting its homework but worse at answering new questions.
Run B was pure LoRA at 1%. Better generalization (validation loss improved from 0.479 to 0.453). But entity tagging nearly disappeared, dropping from 9 tags on test cases to just 2.
| Metric | Base Model | Run A (24% params) | Run B (1% params) |
|---|---|---|---|
| Entity tags (HTN case) | 0 | 3 | 2 |
| Entity tags (DM2 case) | 0 | 6 | 0 |
| Validation loss | - | 0.479 | 0.453 |
Entity tagging requires the model to produce output it has never seen before. The only way to teach new output patterns is to train the layers responsible for generating words. But with only 1,190 examples, that much capacity leads to memorization. I shipped Run A's best checkpoint because entity tagging, even partially working, was more valuable than slightly better generalization without it.
Shipping the App
While fine-tuning was the core ML challenge, shipping an offline medical app required solving several other problems.
Transcription. I chose MLX, Apple's machine learning framework (50MB), over PyTorch (2GB) to run speech recognition natively on the Mac's GPU. Two production bugs shaped the design: the GPU crashes when two AI models try to use it simultaneously (fixed by queuing), and long recordings exceed memory limits (fixed by transcribing in 20-second chunks).
Encryption. Medical data must be encrypted on disk. Envelope encryption uses two keys: a random key that encrypts the data, and the user's password that locks the random key. Changing your password just re-locks the random key, instant regardless of database size.
SQLCipher handles the database encryption using AES-256, the same standard used in banking and government systems. Argon2id hashes the password slowly on purpose so brute-force attacks hit a wall. When the app locks, all keys are wiped from memory.
Speaker diarization. I tried to automatically identify who is speaking, doctor or patient. It wasn't reliable. I removed it entirely. Shipping nothing beats shipping something unreliable in a medical context.
The Thread
Looking back, the pattern is obvious. I kept correcting myself:
- Corrected the voice pipeline (plugged in a language model that was already sitting there)
- Corrected the evaluation rubric (three flaws found and fixed)
- Corrected the model choice (swapped from Gemma 3n to MedGemma)
- Corrected the formatting approach (stopped fighting model pre-training)
- Corrected the approach order (evals before fine-tuning, because you can't fine-tune a model you haven't chosen yet)
None of these corrections came from planning better upfront. They came from looking at what was actually happening and being willing to change direction. The evals didn't just help me pick a model. They revealed that my scoring was flawed, my formatting was fighting the models, and my assumptions about which model was better were wrong.
If I had just picked the first model (Gemma 3n) and started fine-tuning without evaluating first, I'd be polishing the wrong thing right now.
KasaMD runs completely offline, encrypts all patient data, generates 7 types of clinical notes (plus custom templates), and fits in a 4GB download. You can check it out at kasamd.com.